ECDC urges adapting Candida auris lab testing, early control measures
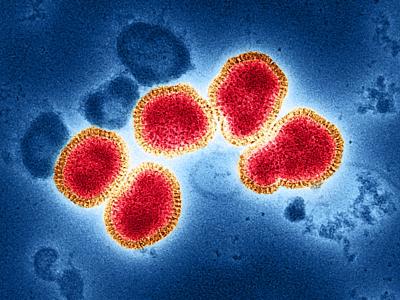
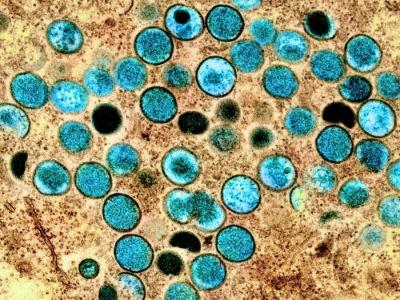
In an updated risk assessment on Candida auris, an emerging difficult-to-treat fungus that can spread in healthcare settings, the European Centre for Disease Prevention and Control (ECDC) said yesterday that the rising number of cases in Europe is concerning because of problems with lab identification and overall lack of awareness.
The antimicrobial-resistant fungus, first reported in Japan, has spread to five continents, and the ECDC said that from 2013 through 2017, 620 cases have been reported in six European countries. The illnesses are often resistant to multiple classes of antifungal medications. The ECDC published an initial risk assessment in December 2016.
In a press release, Dominique Monnet, PharmD, PhD, who leads the ECDC's Antimicrobial Resistance and Healthcare-associated Infections Disease Program, said there's a need to raise awareness in European health facilities, so they can adapt lab testing strategies and implement control measures early enough to stem more hospital outbreaks.
"The occurrence of new outbreaks can be expected. It is therefore of concern that some countries lack national laboratory reference capacity for mycology, or have no information on Candida auris cases available at national level," he said.
Apr 23 ECDC press release
Apr 23 ECDC risk assessment
Antimicrobial resistance evolution tied to regulatory gene
Researchers from Oxford University have uncovered the genetic catalysts behind antibiotic resistance evolution. The work was published yesterday in Nature Ecology & Evolution.
The scientists tested the hypothesis that different bacteria develop resistance at different rates by challenging species of the Pseudomonas genus—bacteria that cause disease in humans, animals, and plants—with the antibiotic ceftazidime.
They found that different species of Pseudomonas developed resistance to ceftazidme at different rates because of the actions of a master regulatory gene, called ampR.
"Species that carry the ampR gene evolve resistance at a higher rate than species that lack this gene," said lead author Craig MacLean, PhD, from Oxford's department of zoology, in a press release. "ampR has this effect because it makes it easier for random mutations to increase the expression of antibiotic resistance genes."
The researchers then blocked the development of ampR in P aeruginosa with an enzyme inhibitor called avibactam. When used with ceftazidime, avibactam helped slow antibiotic resistance, a technique, the authors said, that will need to be replicated in in-vivo experiments.
Apr 23 Nat Ecol Evol study
Apr 23 University of Oxford press release