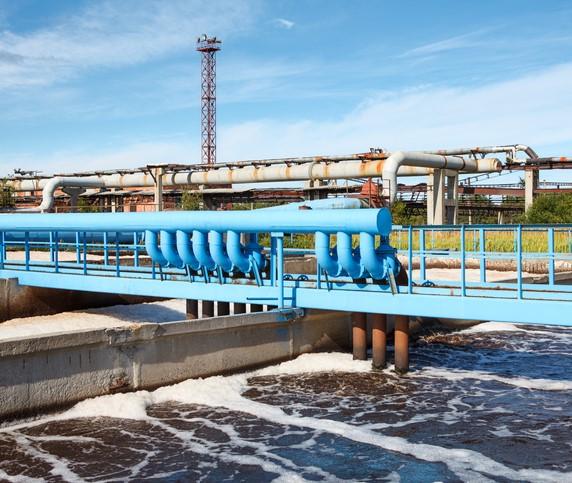
A study today by an international team of scientists has found more evidence of antibiotic resistance in urban wastewater.
In the study, published in Science Advances, the researchers analyzed antibiotic resistance genes in urban wastewater collected at wastewater treatment plants in seven European countries and found that the amount of resistance genes was higher in the wastewater from countries with higher antibiotic use and aligned with the levels of antibiotic resistance found in clinical isolates in the countries.
The results are consistent with the north-to-south pattern that's been observed in studies of antibiotic resistance and consumption in Europe.
The genes that were detected conferred resistance to several classes of antibiotics, including genes that confer multidrug resistance and are of high concern in clinical settings. The study also identified mobile genetic elements that enable bacteria to share and spread resistance.
Relationship between clinical, environmental resistance
The aim of the study was to obtain an initial impression of resistance levels in European urban wastewater, which is considered an important route for the environmental dissemination of antibiotic resistance. The theory is that the content of urban sewage—including human waste, bacteria, and antibiotic residues—might be a good measure of antibiotic resistance in a community because it reflects the contents of the human microbiome.
While surveillance of antibiotic resistance in clinical settings has been conducted for several years and is the primary method for assessing and comparing resistance levels in different countries, data on how much antibiotic resistance exists in the environment is relatively scarce. But several studies in recent months, including a global analysis of resistance genes in sewage from more than 60 countries, have been adding to a growing understanding of the issue.
To select countries for surveillance, the researchers used the most recent data from the European Antimicrobial Resistance Surveillance Network (EARS-Net) as a guide. The EARS-Net data for 2017 shows that clinical antibiotic resistance levels are generally lower in the countries of northern and western Europe and higher among those in the south and the east. Based on these data, the researchers chose 12 wastewater treatment plants in seven countries that reflected that gradient—Portugal, Spain, Cyprus, Ireland, Germany, Norway, and Finland.
Using a quantitative polymerase chain reaction (qPCR) array that targeted 229 antibiotic resistance genes and 25 sequences that involve gene transfer and recombination, the researchers then analyzed the water entering the treatment plants (the influent) and leaving the plants (the effluent), looking for genes found in fecal coliform and enterococci bacteria, which are frequently used as indicators of water quality. Using qPCR allowed the researchers to detect genes that are relevant to human health, which aren't always detected by current metagenomic approaches that focus on the most abundant genes.
Analysis of the influent showed that the relative abundance of antibiotic-resistance genes was much higher in Portugal, Spain, and Cyprus than it was in Germany, Finland, and Norway (influent was not tested in Ireland).
In addition to mirroring the north-to-south gradient of clinical resistance, this clustering also reflected the levels of antibiotic consumption observed in these countries. Antibiotic consumption data show that Finland, Norway, and Germany are routinely among the lowest consumers of human antibiotics in Europe, while Spain, Portugal, and Cyprus are among the highest.
Among the genes found were those conferring resistance to aminoglycosides, beta-lactams, macrolide-lincosamide-streptogramin B, sulfonamides, and tetracyclines. The scientists also detected several genes associated with multidrug-resistant pathogens, including the NDM-1, KPC, VIM, vanA, and MCR-1 genes. In general, the main difference between the two clusters was the much higher abundance of genes conferring resistance to aminoglycosides, sulfonamides, beta-lactams, quinolones, amphenicols and multidrug resistance in the influent in Portugal, Spain, and Cyprus.
Fewer resistance genes in treated wastewater
The analysis of the wastewater effluent aimed to see whether the treatment plants could reduce the levels of resistance genes to similar levels in all countries. And for the most part, treatment was effective at removing many of the resistance genes, including the most clinically concerning multidrug-resistance genes. However, the relative abundance of vancomycin-resistance genes increased in all countries.
Furthermore, the contrast between the high-antibiotic-consumption countries (including Ireland) and the low-consumption countries remained. For most resistance classes, the abundance of resistance genes was still significantly higher in the treated wastewater from Portugal, Spain, and Cyprus, and slightly higher in Ireland, than it was in the effluent from Norway, Finland, and Germany.
This raises the prospect, the researchers say, that the treated water in those countries could be releasing more resistance genes directly into the environment and into irrigation water used in agriculture.
The analysis also found that the amount of resistance gene reduction was higher in the wastewater treatment plants in some of the low-consumption countries, which could be attributed to differences in the way wastewater is treated, and the larger size of treatment plants, in those countries. But the influence of the treatment process was beyond the scope of the study.
Further analysis found that the relative abundance of several resistance genes was significantly correlated with the prevalence of phenotypic resistance in clinical isolates of Escherichia coli, Klebsiella pneumoniae, Pseudomonas aeruginosa, and Staphylococcus aureus—pathogens in which EARS-Net data show a north-to-south resistance gradient.
Other factors in environmental resistance
The authors of the study suggest that factors aside from the levels of antibiotic use and clinical antibiotic resistance might have an influence on the abundance of resistance genes found in the untreated water.
One of those factors is temperature. Average temperatures in the high-consumption countries (with the exception of Ireland) are higher, they note, and a study conducted in 2018 by researchers from MIT found that a temperature increase of 10°C (18°F) across a region coincided with higher resistance levels in E coli, K pneumoniae, and S aureus.
Antibiotic use in livestock could be another factor, as Portugal, Spain, and Cyprus are all high users of antibiotics in food-animal production. Even though antibiotics used in livestock production aren't discharged into urban sewer systems, the authors argue that antibiotic residues from farms may contribute to an increase in the overall antibiotic resistance load in a region.
What this means, they conclude, is that efforts to reduce the spread of antibiotic resistance will likely have to look at all these factors.
"The results not only reinforce the strong relationship between clinical and environmental antibiotic resistance but also signal the importance of taking societal and climate factors (e.g., temperature or precipitation) in design of possible strategies to control antibiotic resistance," they write.
See also:
Mar 27 Sci Adv study
Nov 15, 2018, EARS-Net 2017 report
May 23, 2018, CIDRAP News story "Study finds antibiotic resistance rise tied to hotter temps"