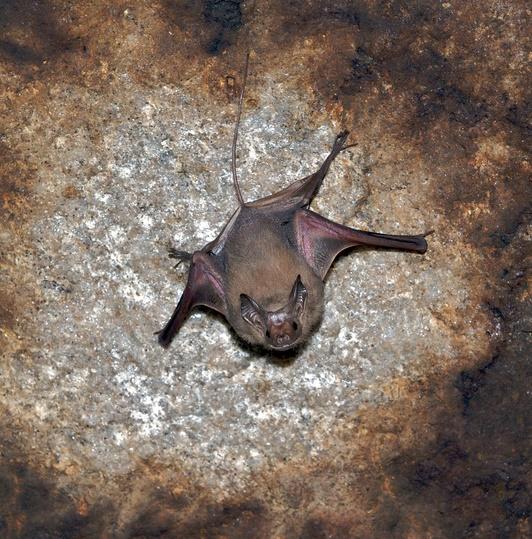
Two new studies shed more light on coronaviruses in bats, one identifying a novel coronavirus similar to MERS-CoV in a bat from Uganda and the other finding wide diversity in China that includes strains similar to the SARS virus.
The new findings add to an expanding list of coronaviruses identified in bats and strengthen the case that the viruses known to cause severe disease in humans originate in bats.
MERS-like virus said to pose no human threat
The study detailing the finding in the Ugandan bat was conducted by a team from the United States and Uganda, part of the US Agency for International Development Emerging Pandemic threats PREDICT project. The researchers published the results yesterday in mBio.
The virus that is similar to MERS-CoV (Middle East respiratory syndrome coronavirus) is called PDF-2180 and was found in a rectal swab of a bat trapped in February 2013. Genetic analysis revealed that it is 87% identical to MERS-CoV and 91% identical to NeoCoV, another coronavirus found in a bat from South Africa. However, differences in part of the spike gene—the part of the virus that invades cells—is significantly different, only 46% identical, from the spike gene of MERS-CoV.
The team also conducted cell-culture studies to see of the virus has the ability to spread to humans. A clone of the virus could reproduce but not enter cells through the human cell receptor typically used by MERS-CoV. It also couldn't establish new infections in Vero cells from monkeys or airway cells from humans.
Simon Anthony, PhD, a study coauthor and assistant professor of epidemiology at the Columbia University Mailman School of Public Health, said in a press release from the college, "In its current form, evolution notwithstanding, this virus is probably not going to be a threat to human health."
SARS similarities in China
In the second study, researchers from China who set out to investigate the diversity and evolution of bat coronaviruses collected 1,067 bats from 2012 to 2015 from 21 species found in five caves in Guizhou, Henan, and Zhejiang provinces. They published their findings yesterday in Virology.
Of 73 coronaviruses they identified, 32 were alphacoronaviruses and 41 were betacoronaviruses. Overall prevalence was 6.84%.
Rhinolophus bat coronaviruses from Guizhou province were closely related to SARS (severe acute respiratory syndrome)-CoV, except for the S gene. The newly identified alphacoronaviruses were genetically diverse, some of which could be novel species, the authors reported.
Phyogenetic analysis to explore how the viruses transmit among the bats revealed clustering by geographic location rather than by species. The testing also showed that interspecies transmission among bats in the same genus was common for both alphacoronaviruses and betacoronaviruses.
Taken together, the findings suggest that high contact rates among bats enable them to acquire and spread coronaviruses, the authors concluded.
See also:
Apr 4 mBio study
Apr 4 Columbia University press release
Apr 4 Virology abstract