Researchers have identified a second sample of human bacteria containing the MCR-1 gene in the United States, according to a study released today.
Italian researchers, meanwhile, said they detected a variant of the MCR-1 gene, which they called MCR-1.2, in Klebsiella pneumoniae.
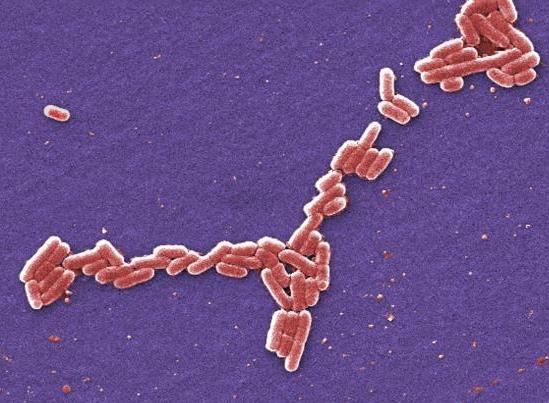
In the US study, which was published in Antimicrobial Agents and Chemotherapy, the researchers reported that they had detected the MCR-1 gene in 19 Escherichia coli isolates from hospitals in 10 countries, including the United States. The MCR-1 gene can make human and animal bacteria resistant to colistin, a powerful antibiotic that is considered a last line of defense against multidrug-resistant bacteria.
The 19 MCR-1 harboring isolates, collected in 2014 and 2015 as part of the SENTRY Antimicrobial Surveillance Program, were found among 13,526 E coli and 7,480 K pneumoniae isolates from 183 hospitals in the Asia-Pacific region, Europe, Latin America, and North America. A total of 390—1.9%—of those isolates showed resistance to colistin. All of the K pneumoniae isolates tested negative for MCR-1.
The isolates were associated with bloodstream infections, skin and skin structure infections, urinary tract infections, respiratory tract infections, and intra-abdominal infections. The US isolate came from a hospital in New York and was collected in May 2015.
Public health officials have voiced concerns about the MCR-1 gene since it was first identified in a strain of E coli recovered from pigs, raw pork meat, and a small number of hospitalized patients in China in November 2015. Because it is located on mobile pieces of DNA called plasmids, MCR-1 has the ability to jump to other types of human and animal bacteria and easily confer colistin resistance to those germs. Since its initial discovery, the gene has been detected in 30 countries and in other types of bacteria.
First human case 6 weeks ago
The finding comes 6 weeks after investigators with the Walter Reed Army Institute of Research announced the first US case of human bacteria carrying the MCR-1 gene. That case involved a 49-year-old Pennsylvania woman with a urinary tract infection caused by E coli.
The authors of the new study said that all 19 isolates were susceptible to several antimicrobial agents, including carbapenems, and are therefore unlikely to cause hard-to-treat infections at this time. But they warned that the prospect of a mobile gene that could encode colistin resistance, particularly to multidrug-resistant bacteria like carbapenem-resistant Enterobacteriaceae (CRE), raises the specter of "superbug" infections that are untreatable.
"The potential dissemination of the mcr-1 among Enterobacteriaceae isolates highlights the importance of active surveillance of emerging resistance genes, including mcr-1 and development of novel antimicrobial agents active against MDR [multidrug-resistant] Gram-negative isolates and CRE," the authors wrote.
MCR-1 variant found in K pneumoniae strain
The MCR-1.2 finding in Italy involved a sample of K pneumoniae from a child with leukemia.
In the study, also published today in Antimicrobial Agents and Chemotherapy, the authors said the structure of the plasmid carrying the gene was similar to that of MCR-1–bearing plasmids previously found in E coli and K pneumonia. They also noted that this is the first time an MCR-type gene has been associated with K pneumoniae ST512, a strain of the bacteria that is resistant to carbapenems.
See also:
Jul 11 Antimicrob Agents Chemother study on MCR-1
Jul 11 Antimicrob Agents Chemother study on MCR-1.2
May 26 CIDRAP News story on first detection of MCR-1 gene in the US