March 30, 2007 (CIDRAP News) – Genetic elements that confer multidrug resistance (MDR) in both plague and foodborne bacteria have a common origin and may represent a significant public health threat, according to a study published Mar 20 in the journal PLoS One (Public Library of Science One).
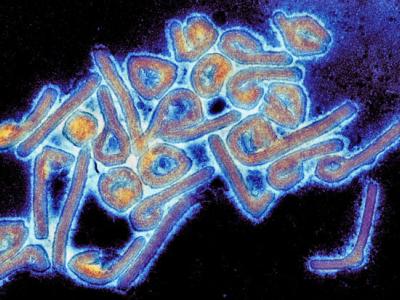
The MDR strain of Yersina pestis (IP275) was identified in 1995 in a single patient in Madagascar who had bubonic plague, the disease that caused the "Black Death" in Europe in the mid 1300s and continues to crop up in small outbreaks today. IP275 exhibited high-level resistance to at least eight drugs used for treating plague, including streptomycin, tetracycline, chloramphenicol, and sulfonamides. An additional strain of MDR plague has since been described in another patient.
In the Madagascar case, antibiotic resistance was found to arise from a self-transmissible plasmid, a small ring of DNA that can move between bacteria. Transmission of plasmids is one way bacteria acquire new genetic material, including genes that confer resistance to drugs. Increased drug resistance can also arise from indiscriminate use of antimicrobials in people and in agriculture, according to many infectious disease experts.
In the PLoS article, an international team of researchers report finding common elements in the sequences of MDR plasmids from plague, a Salmonella species, and a related Yersinia species, an unexpected result that heightens concern about the spread of antibiotic resistance and re-emergence of plague. The first author of the report is Timothy J. Welch, an aquaculture researcher with the US Department of Agriculture.
Related MDR plasmids in multiple species
The investigators analyzed and compared the sequences of large, nearly identical MDR plasmids from Y pestis strain IP275; Salmonella enterica serotype Newport, a foodborne pathogen; and Y ruckeri, a fish pathogen. Comparisons of plasmid gene sequences demonstrated that they were closely related and had a common origin, the report says.
Sequence analysis of plasmids revealed nearly identical "backbones," consisting of 135 genes located at similar positions, and resistance genes for multiple antimicrobial agents. The common backbone also contained a gene conferring resistance to sulfonamides and four locations that had inserted antimicrobial resistance genes. All plasmids also had another area that can act as a "hot spot" for insertion of foreign DNA.
The presence of closely related antibiotic-resistance plasmids in different species of bacteria is not a new finding, but this example suggests that all these organisms had recently acquired plasmids from a common source, the authors say. They base this belief on the presence of resistance to sulfonamides, drugs first introduced into clinical use in the 1930s. The authors acknowledge that the exact timing of the plasmid introduction remains unknown, however. They suggest that one possible mechanism for plasmid transfer to Y pestis may have been coinfection of a mammalian host or in the midgut of the flea.
"The fact that we found a plasmid usually found in Salmonella in Y pestis is a big problem," said Jacques Ravel of The Institute for Genomic Research (TIGR), senior author of the study. "It also raises a question about how this happened, how it went from one to the other." Ravel was quoted in a news release from TIGR.
MDR plasmids found in foodborne pathogens
Although MDR plasmids in Y pestis and Y ruckeri are rare, antimicrobial resistance in other bacteria such as foodborne Salmonella is more common. Surveillance data from many countries indicate that the incidence of MDR Salmonella has been increasing, and the authors sought to determine the type of MDR resistance in foodborne pathogens and compare it with MDR resistance found in plague.
The research team used gene sequencing techniques to analyze the occurrence and distribution of the common plasmid background in three sets of samples: 125 MDR Salmonella strains recovered from retail meats from 2002 to 2005 through the National Antimicrobial Resistance Monitoring System (NARMS), a small collection of E coli strains recovered from food samples, and Klebsiella isolates from ground turkey meat from Iowa.
The authors detected the common MDR plasmid backbone in multiple Salmonella serotypes, nine samples of Klebsiella from ground turkey, and E coli isolated from a calf and from ground turkey. The discovery of these MDR plasmids in evolutionarily distinct bacteria indicates recent genetic exchange, either directly between species or through bacterial intermediates, according to the authors. They suggest that the overlapping ranges of these organisms may have aided past transmission, and perhaps may facilitate future transmission between organisms, a potentially dangerous occurrence.
Resistance transferred experimentally
The researchers also conducted transfer experiments with the plasmids found in 70 MDR-positive foodborne organisms to assess the potential for interspecies transfer. The investigators were able to transfer related-resistance plasmids from 30 of the foodborne pathogens to Y ruckeri, a plague-related pathogen. These experiments demonstrate a potential for Y pestis and other animal pathogens to acquire resistance, although the authors acknowledge that other factors such as additional genes and host protective systems may also influence plasmid transfer.
The potential for transfer should not be underestimated, however, the authors state. Elisabeth Carniel of the Pasteur Institute in Paris, a co-author of the report, said in the press release, "When we identified the first Y pestis strain resistant to multiple antibiotics, we warned that if this type of strain spreads or emerges again, it would pose a serious health problem. The discovery that the multidrug resistance plasmid acquired by the plague bacillus is widespread in the environmental bacteria reinforces this warning."
The authors suggest that antimicrobial resistance monitoring should be expanded, especially in areas such as Africa, Asia, and the southwestern United States, where both Y pestis and MDR Salmonella are found and are likely to come into direct contact. The investigators also say their methodology can provide the means to monitor such plasmids in pathogens recovered from diverse environments.
Antibiotic overuse may promote resistance transfer
Olaf Schneewind, MD, PhD, a microbiologist who has studied how plague affects the immune system, commented that the discovery of transfer of antibiotic resistance between bacterial species is not new, but agreed that the authors' findings suggest a possible public health threat. He told CIDRAP News that overuse of antibiotics, particularly in agriculture, increases the likelihood of transfer of drug resistance to virulent organisms.
"Widespread use of antimicrobials leads to drug resistance in organisms we would not usually expect, and that creates a potential public health risk," said Schneewind, chairman of the department of microbiology in the biological sciences division at the University of Chicago. Schneewind has examined mechanisms and strategies of pathogenic bacteria, including plague, and plague vaccine candidates.
Schneewind noted that as a zoonotic disease, plague exists in the background in many areas, needing only the right circumstances to cause outbreaks. An outbreak of MDR plague in an area well-connected to the rest of the world could pose a serious public health problem, he said.
Welch TJ, Fricke WF, McDermott PF, et al. Multiple antimicrobial resistance in plague: an emerging public health threat. PLoS One 2007 Mar 20;2(3):e309 [Full text]
See also:
Mar 20 news release from TIGR
http://www.jcvi.org/cms/press/press-releases/full-text/article/antibiotic-resistance-in-plague/
Galimand M, Carniel E, Courvalin P. Resistance of Yersinia pestis to antimicrobial agents. Antimicrob Agents Chemother 2006 Oct;50(10):3233-36 [Full text]